Biomedical Imaging and Informatics
Department of Computer Science and Engineering
University of Washington
Biomedical Imaging and Informatics
The Biomedical
and Health Informatics Program at the University of
Washington is a graduate program in the UW Medical School. Our
research group collaborates with and includes students and faculty
from this program. Here are some of our collaborative projects.
CT Image Registration, Segmentation, and Retrieval
Jiun-Hung Chen is working on construction of 3D models from
image data. His current work concentrates on finding
point correspondences between 3D medical entities such
as abdominal organs.
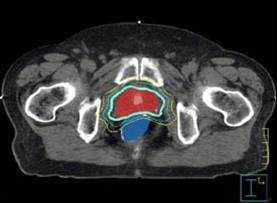
Chia-Chi Teng's 2007 dissertation describes a new methodology for
obtaining contours of lymph node regions in CT scans of cancer
patients. The method first identifies the most similar reference
patients using a similarity measure based on landmarks that can
be reliably extracted from the images. It then uses a constrained
optimization procedure to find the best deformable mapping from
the new patient to the reference patient. The mapping is used to
map the predrawn contours on the reference patient to the CT
scan of the new patient, so that the lymph node regions can be
reliably found and the radiation treatment accordingly planned.
Publications:
- J. H. Chen and L. G. Shapiro,
"Groupwise Pose Normalization for Craniofacial Applications,"
IEEE Workshop on Applications of Computer Vision, January 2011.
- J.H. Chen, K. Zheng, and L. G.
Shapiro, "3D Point Correspondence by Minimum Description Length in Feature Space," European Conference on Computer Vision,
2010.
- J. H. Chen and L. G. Shapiro,
"3D Point Correspondence by Minimum Description Length with
2DPCA," International Conference of the IEEE Engineering in
Medicine and Biology Society (EMBC'09), 2009.
- J. H. Chen and L. G. Shapiro,
"PCA vs. Tensor-Based Dimension Reduction Methods: An
Empirical Comparison on Active Shape Models of Organs,"
International Conference of the IEEE Engineering in Medicine
and Biology Society (EMBC'09), 2009.
- J. H. Chen and L. G. Shapiro, "Medical Image Segmentation via min s-t Cuts with Side Constraints," International Conference on
Pattern Recognition , 2008.
- C. Teng, L. G. Shapiro, and I. Kalet, ``Head and Neck Cancer Patient
Similarity Based on Anatomical Structural Geometry,'' IEEE
International Symposium on Biomedical Imaging,
2007.
- C. Teng, L. G. Shapiro, and I. Kalet,
"Head and Neck Lymph Node Region Delineation
Using a Hybrid Image Registration Method,"
Proc. IEEE International Symposium on Biomedical Imaging,
April 2006, pp. 462-465.
- C. Teng, L.G. Shapiro, and I. Kalet,
"Automatic Segmentation of Neck CT Images,"
Proc. IEEE International Symposium on Computer-Based Medical Systems (CMBS), June 2006.
- C. Teng, M. M. Austin-Seymour, J. Barker, I. J. Kalet, L. G. Shapiro, M. Whipple,
"Head and Neck Lymph Node Region Delineation with 3-D CT Image Registration,"
Proceedings American Medical
Informatics Association Annual Symposium, 2002, pp. 767-771.
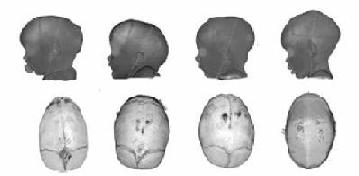
3D Skull Shape Analysis and Retrieval
Salvador Ruiz Correa and Jill Lin have worked on automatic classification
and retrieval of skull images for craniofacial applications
Jill Lin's 2006 dissertation described a new methodology for
describing 3D shapes, including skulls and feet, in terms of
symbolic shape descriptors that could then be used to classify
shapes or to retrieve similar ones.
Publications:
-
S. Ruiz-Correa, D. Gatica-Perez,
H. J. Lin, L. G. Shapiro, and R.W. Sze,
"Bayesian Hierarchical Model for Classifying Craniofacial
Malformations from CT Imaging," IEEE EMBS Annual International Conference , 2008.
-
H. J. Lin, S. Ruiz-Correa, L. G. Shapiro,
M. L. Cunningham, M. Speltz, and W. R. Sze,
"Predicting Neuropsychological Development from Skull Imaging,"
28th IEEE EMBS Annual International Conference, 2006.
-
S. Ruiz-Correa, R. W. Sze, J. R. Starr,
H. J. Lin, M. L. Speltz, M. L. Cunningham, A. V. Hing,
"New Scaphocephaly Severity Indices of Sagittal Craniosynostosis:
A Comparative Study with Cranial Index Quantifications.
-
H. J. Lin, S. Ruiz-Correa, L. G. Shapiro, M. L. Cunningham, and W. R. Sze,
"Efficient Symbolic Signatures for Classifying Craniosynostosis Skull
Deformities,"
Proceedings of the Workshop on Computer Vision for Biomedical
Image Applications, Oct. 2005, pp. 302-313.
Ulstrasound Image Analysis
Publications:
Anatomy Ontologies
The Foundational Model of Anatomy is a large reference ontology for
the human body developed by Dr. Cornelius Rosse and his colleagues.
We have participated in the development of this model and constructed
several query systems that employ graphical user interfaces to access
its data.
The Emily graphical query inteface allows the user to query
the FMA for relational information about the more than 80,000
anatomical entities represented in the ontology.
The CAIS system represents the 2006 Ph.D. dissertation work
of Ravensara Travillian who developed the Structural Difference Method
(SDM), a method for comparing pairs of species motivated by graph
matching. The CAIS graphical user interface allows users to compare
2 species (human, mouse, rat) with respect to the entities and
relationships found in the FMA.
Publications:
- Y. Chen, H.Lu, L. Shapiro, R. S. Travillian, L. Li, "An Approach to Semantic Query Expansion System Based on Hepatitis
Ontology," Journal of Biological Research-Thessaloniki, 23(Suppl 1):S11, 201
6.
-
R. S. Travillian, K. Diatchka, T. K. Judge, K. Wilamowska, L. G. Shapiro, "An Ontology-Based Comparative Anatomy Information System," Artificial Intelligence in Medicine, Vol. 51, 2011, pp. 1-15.
-
G. Yngve, J.F. Brinkely, D. Cook, L. G. Shapiro,
"A Model Browser for Biosimulation",
AMIA 2007 Annual Symposium 2007
-
R. S. Travillian, K. Diatchka, T. K. Judge, K. Wilamowska, and L. G. Shapiro, "A Graphical User Interface for a Comparative
Anatomy Information System: Design, Implementation and Usage Scenarios",
AMIA 2006 Annual Symposium 2006
-
R. S. Travillian, J. H. Gennari, and L. G. Shapiro,
"Of Mice and Men: Design of a Comparative Anatomy Information System",
Proceedings of the Annual Symposium of the American Medical
Informatics Association, October 2005.
-
L. G. Shapiro, E. Chung, L. Detwiler,
J. Mejino, Jr., A. Agoncillo, J. F. Brinkley, and C. Rosse,
"Processes and Problems in the Formative Evaluation
of an Interface to the Foundational Model of Anatomy Knowledge Base,"
Journal of the American Medical Informatics Association,
Vol. 12, 2005, pp. 35-46.
-
L. T. Detwiler, E. Chung, A. Li,
J. L. V. Mejino Jr., A. V. Agoncillo, J. F. Brinkley,
C. Rosse and L. G. Shapiro, "A Relation-Centric Query Engine
for the Foundational Model of Anatomy," Proceedings of MedInfo ,
2004.
-
R. S. Travillian, C. Rosse, L. G. Shapiro,
"An Approach to the Anatomical Correlation of Species,"
Proceedings of the AMIA Symposium, 2003, pp. 669-673.
-
P. J. Neal, L. G. Shapiro and C. Rosse,
"The Digital Anatomist Structural Abstraction: a Scheme
for the Spatial Description of Anatomical Entities'',
Proceedings 1998 American Medical Informatics
Association Annual Symposium, Lake Buena Vista, FL, 7-11 November 1998.
Demos: